Typically, you’ll want to use a validation set to determine an optimal threshold as it is often not .5 (which is equivalent to argmax). Subsequently, use this threshold on the the “_prob” image to generate a binary image.This blog posts explains how to train a deep learning lymphocyte detector in accordance with our paper “Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases”.
Use Case 2: Epithelium Segmentation
This blog posts explains how to train a deep learning epithelium segmentation classifier in accordance with our paper “Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases”.
Use Case 1: Nuclei Segmentation
This blog posts explains how to train a deep learning nuclear segmentation classifier in accordance with our paper “Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases”.
Exporting from Matlab To PowerPoint
Reviewing the results of an image based experiment, across many images, can be annoying in matlab. Too much clicking!
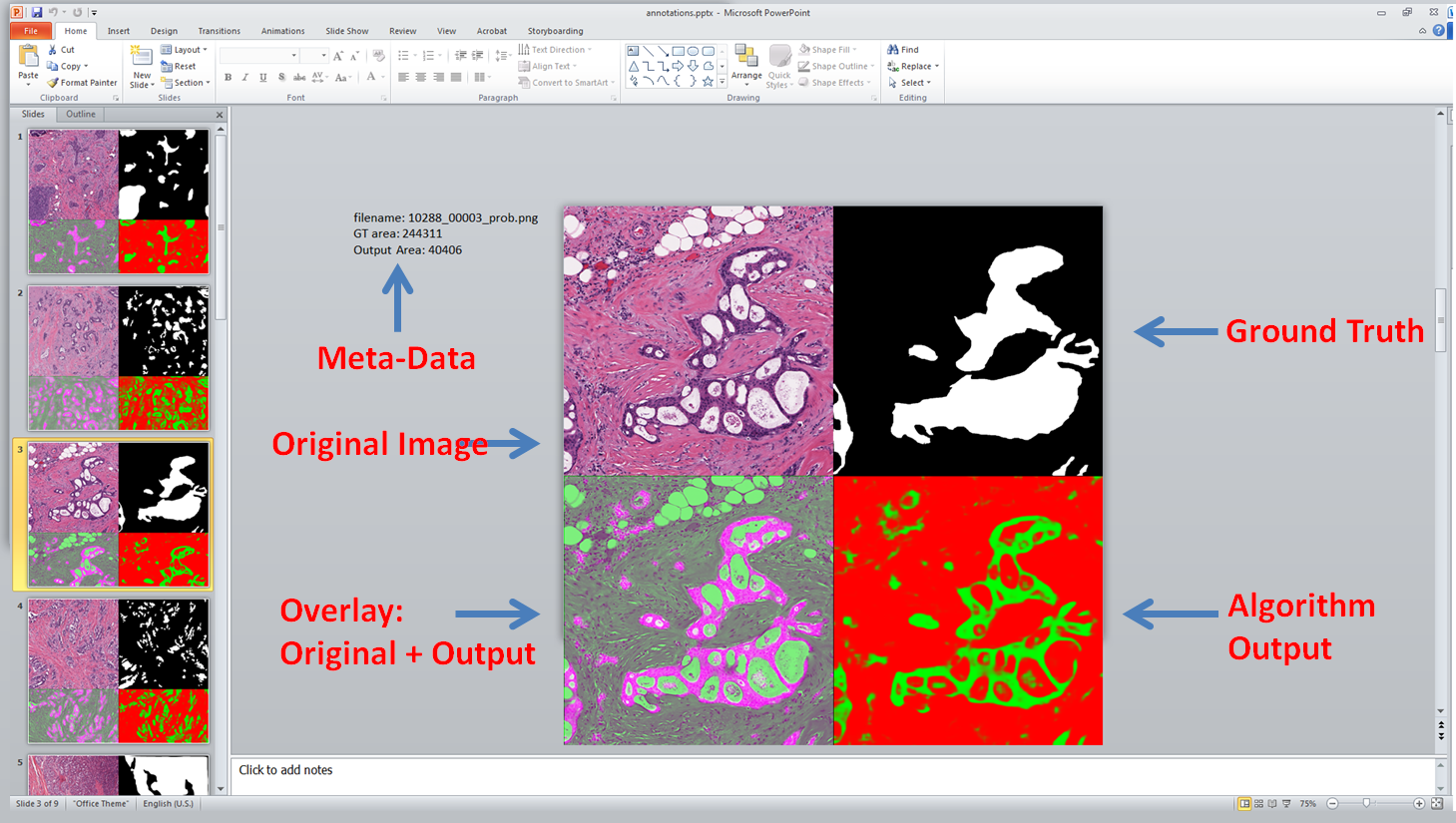
I’ve recently started using PowerPoint to view many of my results. This blog posts discuss how using the free export to PowerPoint toolbox it is possible to create a slide desk with all relevant information for easier viewing. It looks like this:
Installing Caffe on the Ohio Super Computing (OSC) Ruby Cluster
One of the perks of working at Case Western Reserve is that we often qualify for access to cutting edge resource and special projects. In this case, since our digital histology deep learning work requires a large number of GPUs to analyze thousands of patients, we were granted access to the OSC Ruby cluster, which has 20 NVIDIA Tesla K40 GPUs. Since the cluster has only recently been setup, there was some leg work required on our end to get Caffe fully up and running, without root access, which we’ll document here.
Continue reading Installing Caffe on the Ohio Super Computing (OSC) Ruby Cluster
Exporting Bisque Gobjects/Annotations as Binary Masks into Matlab
Annotations which stay solely in Bisque aren’t incredibly useful. One of the main reasons for marking up images is to be able to use those annotations for some other purpose, such as training classifiers, computing metrics and features. In this post, we show how to take the annotations from bisque via REST, and convert them into binary masks in matlab.
Continue reading Exporting Bisque Gobjects/Annotations as Binary Masks into Matlab
Created Bisque XML from Matlab Binary Masks
Once we have images uploaded to Bisque, we might want to be able to overlay annotations on top of them so that users can interact with them. This post quickly goes through the conversation of a binary mask, in matlab, into an XML which can later be imported into Bisque.
Continue reading Created Bisque XML from Matlab Binary Masks
Uploading Bisque XML via Python
Assuming we have the necessary files on our Bisque server (perhaps uploaded with our script), and we have a set of bisque compliant XML annotations (perhaps generated with our script), we would like to upload them to the Bisque server so that they can be evaluated or modified. That is what this post is about 🙂
Uploading Files to Bisque via Python
In this blog post, we discuss how to quickly upload a dataset to Bisque using python. The next blog post will then talk about how to convert binary masks (made in matlab) to annotations usable by Bisque for validation or modification.
Import Annotations from Matlab into BigTiff XML (Ventana)
In the previous post we discussed how to export annotations from a Ventana Image Viewer program and create binary masks. Now we explain how to do the opposite and import the mask back into Image Viewer.
Continue reading Import Annotations from Matlab into BigTiff XML (Ventana)

