This blog posts explains how to train a deep learning epithelium segmentation classifier in accordance with our paper “Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases”.
Please note that there has been an update to the overall tutorial pipeline, which is discussed in full here.
This text assumes that Caffe is already installed and running. For guidance on that you can reference this blog post which describes how to install it in an HPC environment (and can easily be adopted for local linux distributions).
Preface
As mentioned in the paper, the goal of this project was to create a single deep learning approach which worked well across many different digital pathology tasks. On the other hand, each tutorial is intended to be able to stand on its own, so there is a large amount of overlapping material between Use Cases.
If you’ve already read the Use Case 1: Nuclei Segmentation, in this tutorial you can focus on “Step 1: Patch Extraction”, which contains all the differences from the nuclei tutorial.
Background
The identification of epithelium and stroma regions is important since regions of cancer are typically manifested in the epithelium. Additionally recent work by Beck et al. [31] suggest that histologic patterns within the stroma might be critical in predicting overall survival and outcome in breast cancer patients. Thus from the perspective of developing algorithms for predicting prognosis of disease, the epithelium-stroma separation becomes critical.
Given that our AlexNet approach constrains input data to a 32 x 32 window, we need to appropriately scale the task to fit into this context. The general principal employed is that a human expert should be able to make an educated decision based solely on the context present in the patch supplied to the DL network. What this fundamentally implies is that we must a priori select an appropriate magnification from which to extract the patches and perform the testing. In this particular case, we down sample each image to have an apparent magnification of 10x (i.e., a 50% reduction) so that sufficient context is available for use with the network. Networks which accept larger patch sizes could thus potentially use higher magnifications, at the cost of longer training times, if necessary.
Overview
We break down this approach into 5 steps:
Step 1: Patch Extraction (Matlab): extract patches from all images of both the positive and negative classes
Step 2: Cross-Validation Creation (Matlab): at the patient level, split the patches into a 5-fold training and testing sets
Step 3: Database Creation (Bash): using the patches and training lists created in the previous steps, create 5 sets of leveldb training and testing databases, with mean files, for high performance DL training.
Step 4: Training of DL classifier (Bash): Slightly alter the 2 prototxt files used by Caffe, the solver and the architecture to point to the correct file locations. Use these to train the classifier.
Step 5: Generating Output on Test Images (Python): Use final model to generate the output
There are, of course, other ways of implementing a pipeline like this (e.g., use Matlab to directly create a leveldb, or skip the leveldb entirely, and use the images directly for training) . I’ve found using the above pipeline fits easiest into the tools that are available inside of Caffe and Matlab, and thus requires the less maintenance and reduces complexity for less experienced users. If you have a suggested improvement, I’d love to hear it!
Dataset Description
The dataset consist of 42 images ER+ BCa images scanned at 20x. Each image is 1,000 x 1,000. The epithelium region was manually annotated across all images by an expert pathologist.
The format of the files is:
8918_00007.tif: original H&E image
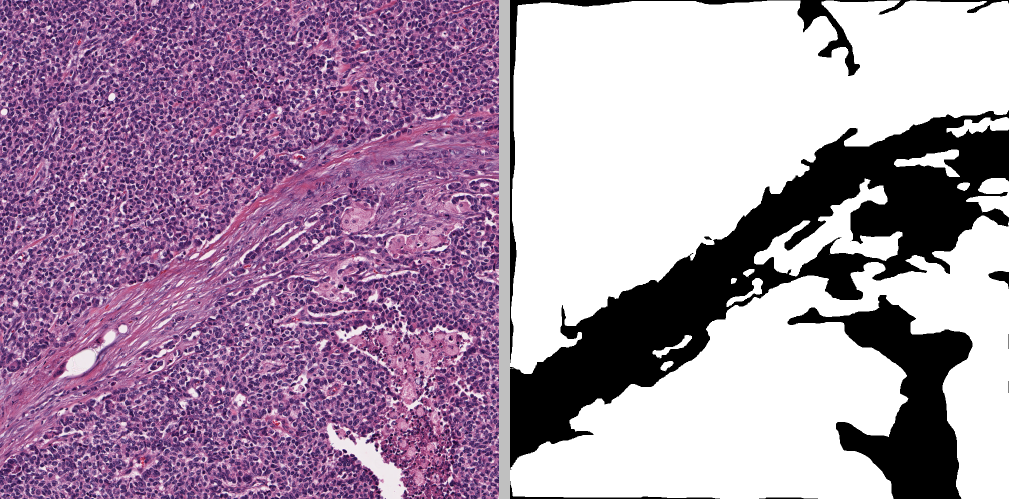
8918_00007_mask.png: mask of the same size, where white pixels indicate epithelium
Each image is prefaced by a code (e.g., “8918”) to the left of the first underscore (_), which coincides with a unique patient number. In this dataset, no patients have more than 1 image associated with them (43 patients = 43 images).
The data and the associated masks are located here (336M).
Examples of these images and their annotates can be seen below
Step 1: Patch Extraction (Matlab)
We refer to step1_make_patches.m, which is fully commented.
A high level understanding is provided here:
- Break the images into groups by patient ID number (just in case later on we add additional images per patient).
- For each image, load the image and its annotation and downsize both of them by 50% (10x apparent magnification), create an edge mask, and a complement to the annotated image (to create a negative class). This task tends to have a lot of background, so we apply a gray level threshold to remove all “white” background regions.
 Original Image |
 Epithelium Annotation |
 Non-Epi (Stroma) Mask |
 Epithelium Edge Mask |
- From each of these, randomly crop patches, save them to disk. At the same time, we maintain a “patient_struct”, which contains all of the file names which have been written to disk.
The file name format for each patch is as follows:
u_v_w_x_y_z.png — > example 10256_1_0_e_1_0.png
Where u is the patient ID, v is the image number of the patient, w is the class (0 (non-epithelium) or 1 (epithelium)), x is the type of patch (“e” is edge, “b” is background (i.e., stroma), “p” is positive), y is the number of the patch (1 to 5000), and z is rotation (0 or 90 in this case).
Step 2: Cross-Validation Creation (Matlab)
Now that we have all of the patches written to disk, and we have all of the file names saved into patient_struct, we want to split them into a cross fold validation scheme. We use step2_make_training_lists.m for this, which is fully commented.
In this code, we use a 5-fold validation, which is split at the patient level. Again, remember splitting at the image level is unacceptable if multiple images can come from the same patient!
For each fold, we create 4 text files. Using fold 1 as an example:
train_w32_parent_1.txt: This contains a list of the patient IDs which have been used as part of this fold’s training set. This is similar to test_w32_parent_1.txt, which contains the patients used for the test set. An example of the file content is:
10256
10260
10261
10262
train_w32_1.txt: contains the filenames of the patches which should go into the training set (and test set when using test_w32_1.txt). The file format is [filename] [tab] [class]. Where class is either 0 (non-epithelium) or 1 (epithelium). An example of the file content is:
10256_1_0_e_1_0.png 0
10256_1_0_e_1_90.png 0
10256_1_0_e_2_0.png 0
10256_1_0_e_2_90.png 0
10256_1_0_e_3_0.png 0
10256_1_0_e_3_90.png 0
All done with the Matlab component!
Step 3: Database Creation (Bash)
Now that we have both the patches saved to disk, and training and testing lists split into a 5-fold validation cohort, we need to get the data ready for consumption by Caffe. It is possible, at this point, to use an Image layer in Caffe and skip this step, but it comes with 2 caveats, (a) you need to make your own mean-file and ensure it is in the correct format and (b) an image layer can is not designed for high throughput. Also, having 100k+ files in a single directory can bring the system to its knees in many cases (for example, “ls”, “rm”, etc), so it’s a bit more handy to compress them all in to 10 databases (1 training and 1 testing for 5 folds), and use Caffe’s tool to compute the mean-file.
For this purpose, we use this bash file: step3_make_dbs.sh
We run it in the “subs” directory (“./” in these commands), which contains all of the patches. As well, we assume the training lists are in “../”, the directory above it.
Here we’ll briefly discuss the general idea of the commands, while the script has additional functionality (computes everything in parallel for example).
Creating Databases
We use the caffe supplied convert_imageset tool to create the databases using this command:
~/caffe/build/tools/convert_imageset -shuffle -backend leveldb ./ DB_train_1
We first tell it that we want to shuffle the lists, this is very important. Our lists are in patient and class order, making them unsuitable for stochastic gradient descent. Since the database stores files, as supplied, sequentially, we need to permute the lists. Either we can do it manually (e.g., use sort –random) , or we can just let Caffe do it 🙂
We specify that we want to use a leveldb backend instead of a lmdb backend. My experiments have shown that leveldb can actually compress data much better without the consequence of a large amount of computational overhead, so we choose to use it.
Then we supply the directory with the patches, supply the training list, and tell it where to save the database. We do this similarly for the test set.
Creating mean file
To zero the data, we compute mean file, which is the mean value of a pixel as seen through all the patches of the training set. During training/testing time, this mean value is subtracted from the pixel to roughly “zero” the data, improving the efficiency of the DL algorithm.
Since we used a levelDB database to hold our patches, this is a straight forward process:
~/caffe/build/tools/compute_image_mean DB_train_1 DB_train_w32_1.binaryproto -backend leveldb
Supply it the name of the database to use, the mean filename to use as output and specify that we used a leveldb backend. That’s it!
Step 4: Training of DL classifier (Bash)
Setup files
Now that we have the databases, and the associated mean-files, we can use Caffe to train a model.
There are two files which need to be slightly altered, as discussed below:
BASE-alexnet_solver.prototxt: This file describes various learning parameters (iterations, learning method (Adagrad) etc).
On lines 1 and 10 change: “%(kfoldi)d” to be the number of the fold for training (1,2,3,4,5).
On line 2: change “%(numiter)d” to number_test_samples/128. This is to have Caffe iterate through the entire test database. Its easy to figure out how many test samples there are using:
“wc –l test_w32_1.txt”
BASE-alexnet_traing_32w_db.prototxt: This file defines the architecture.
We only need to change lines 8, 12, 24, and 28 to point to the correct fold (again, replace “%(kfoldi)d” with the desired integer). That’s it!
Note, these files assume that the prototxts are stored in a directory called ./model and that the DB files and mean files are stored in the directory above (../). You can of course use absolute file path names when in doubt.
In our case, we had access to a high performance computing cluster, so we used a python script (step4_submit_jobs.py) to submit all 5 folds to be trained at the same time. This script automatically does all of the above work, but you need to provide the working directory on line 11. I use this (BASE-qsub.pbs) PBS script to request resources from our Torque scheduler, which is easily adaptable to other HPC environments.
Initiate training
If you’ve used the HPC script above, things should already be queued for training. Otherwise, you can start the training simply by saying:
~/caffe/build/tools/caffe train –solver=1-alexnet_solver_ada.prototxt
In the directory which has the prototxt files. That’s it! Now wait until it finishes (600,000) iterations. 🙂
Step 5: Generating Output on Test Images (Python)
At this point, you should have a model available, to generate some output images. Don’t worry, if you don’t, you can use mine.
Here is a python script, to generate the test output for the associated k-fold (step5_create_output_images_kfold.py).
It takes 2 command line arguments, base directory and the fold. Make sure to edit line 88 to apply the appropriate scaling; in this case reduce 20x images to 10x images by resizing by 1/2.
The base directory is expected to contain:
BASE/images: a directory which contains the tif images for output generation
BASE/models: a directory which holds the 5 models (1 for each fold)
BASE/test_w32_parent_X.txt: the list of parent IDs to use in creating the output for fold X=1,2,3,4,5, created in step 2
BASE/DB_train_w32_X.binaryproto: the binary mean file for fold X=1,2,3,4,5, created in step 3
It generates 2 output images for each input. A “_class” image and a “_prob” image. The “_prob” image is a 3 channel image which contains the likelihood that a particular pixel belongs to the class. In this case, the Red channel represents the likliehood that a pixel belongs to the non-epithelium class, and the green channel represents the likelihood that a pixel belongs to the epithelium class. The two channels sum to 1. The “_class” image is a binary image using the argmax of the “_probs image”.
Notice that the output masks are 500 x 500, or 1/2 the size of the input masks. This is very important, we trained the classifier at 10x, so we need to test it at 10x. Since the images are all taken at 20x, we down-sample them to obtain the correct apparent magnification .
 |
 |
 |
| Input Image | _class image | _prob image |
Typically, you’ll want to use a validation set to determine an optimal threshold as it is often not .5 (which is equivalent to argmax). Subsequently, use this threshold on the the “_prob” image to generate a binary image.
Final Notes
Efficiency in Patch Generation
Writing a large number of small, individual files to a harddrive (even SSD) is likely going to take a very long time. Thus for Step 1 & Step 2, I typically employ a ram disk to drastically speed up the processes. Regardless, make sure Matlab does not have the output directory in its search path, otherwise it will likely crash (or come to a halt), while trying to update its internal list of available files.
As well, using a Matlab Pool (matlabpool open), opens numerous workers which also greatly speed up the operation and is recommended as well.
Efficiency in Output Generation
It most likely will take a long time to apply a the classifier pixel wise to an image to generate the output. In actuality, there are many ways to speed up this process. The easiest way in this case is to simply use a larger stride such that you compute every 2nd or 3rd pixel since epithelium segmentation doesn’t require nearly as much precision, as say, nuclei segmentation.
Keep an eye out for future posts where we delve deeper into this and provide the code which we use!
Magnification
It is very important to use the model on images of the same magnification as the training magnification. This is to say, if your patches are extracted at 10x, then the test images need to be done at 10x as well.
Code is available here
Data is available here (336M)



Thank you very much for posting this tutorial. I have a question regarding Step 5 of the process. I am getting the following error whenever I try to run the python script:
Traceback (most recent call last):
File “step5_create_output_images_kfold.py”, line 104, in
outputimage_probs[rowi,hwsize+1:image.shape[1]-hwsize,0:2]=prediction #save the results to our output images
ValueError: could not broadcast input array from shape (200,3) into shape (999,2)
Do you possibly have any idea how I may be able to address this?
I would guess that you changed your stride variable from 1 to 5? That would explain why your output is 200 columns wide, but the original image is 1000 columns wide.
that being the case, i guess there is an error in my code and the line should read:
outputimage_probs[rowi,hwsize+1:image.shape[1]-hwsize:stride,0:2]=prediction #save the results to our output images
to make appropriate stride sized steps.
In fact, I don’t use any of that code anymore and would recommend you look at these two posts: updated pipeline and efficient output generation, which contain all the latest and greatest stuff we’ve made public.
Thank you very much. I will be sure to look at the newer posts.
If possible, however, I wanted to ask one more question about this pipeline.
Would you mind explaining what you meant by the following lines: “Typically, you’ll want to use a validation set to determine an optimal threshold as it is often not .5 (which is equivalent to argmax). Subsequently, use this threshold on the the “_prob” image to generate a binary image.”
The prob image I am generating is accurate, but the binary mask is not.
a softmax classifier simply selects whichever class has the highest probability. in the extreme case, this implies that if there are two cases and one has 50.00001% and the other has 49.99999%, the classifier will select the first case. while this is useful for training, in practice, the sensitivity of the DL classifier is usually at a slightly different boundary. maybe everything > 60% should be considered as positive (reducing false positives) or maybe everything >40% (reducing false negatives). each test set is slightly different, and given that the classifier is trained starting from a random initialization, each classifier is slightly different as well. the way to find the optimal operating point is to use 3 sets of data. you train on the training data, then use the validation data to the find the threshold which maximizes whatever your function is (e.g., accuracy, f-score, etc), and then use that threshold on the test set to generate the final data. note, in order to do this, one needs to have labels for the validation data, but not for the test data (which would be cheating).
are the whole slides from which these patches were extracted available for download?
Unfortunately, they aren’t our resource so we can’t release directly, but we’re in progress of obtaining official approval for a release
Hi! I am new to linux. During step 4 It gave me an error “: 1: : qsub: not found”. Kindly help me to resolve this issue.Thanks
That is a job scheduler for a high performance computing cluster. If you’re running it on a local machine (with a GPU) you can dissect the commands and run them directly
Can you give me a example of command without cluster?
how about you guess and i’ll tell you if you’re right? this is a good learning experience!
Thank you for this valuable tutorial.
In the data set used for epithelium segmentation, we have just 42 mask image. Can you send ground-truth for all images.
I need more data and label.
Thank you
unfortunately, this is all that i have, sorry