Aperio scanners generate a semi-proprietary file format called SVS. At its heart, SVS files are really a multi-page tiff file storing a pyramid of smaller tiff files of the original image. We’ll look at those here using a SVS file provided by the TCGA (http://cancergenome.nih.gov/) breast cancer cohort:
TCGA-A1-A0SD-01Z-00-DX1.DB17BFA9-D951-42A8-91D2-F4C2EBC6EB9F.svs
First we’ll open it in Aperio ImageScope to see what we’re looking at:
We see already 4 different versions of the same image, a high-level view, a low level view, a thumbnail and a “working area”. The working area allows us to scroll in and out of regions of interest (ROI).
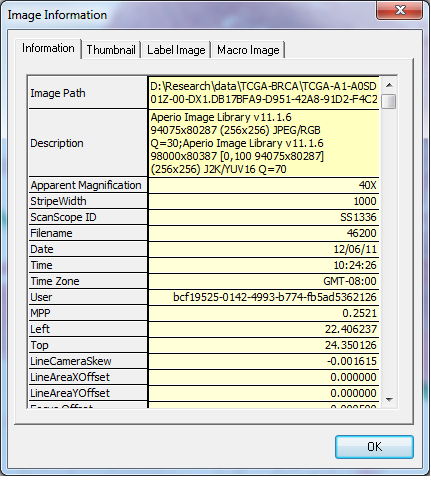
If we use Image->Information, we can see some additional information:
Which tells us the apparent magnification which the specimen was scanned at (40X) and microns-per-pixel (MPP), which are important to know when working with images from different sources.
Additionally, we can take a peek at the various layers in the Pyramid:
We can see that at the base, the image is 94k x 80k. This is the uncompressed original image scanned at the apparent magnification (40x). The rest of them are further down-sampled versions of the original. We can see at Level 1, we have a ratio of 4:1, meaning that the image stored there is 25% the side of the original image.
The main reason for this is simple, loading the entire image is very time consuming, if not impossible based on memory constraints. If we look at the lowest resolution necessary to fill every pixel on the screen, essentially the user can’t tell the difference. Each point between the different pyramids is linked together based on the ratios, so knowing the location at one level can easily be mapped into a higher level (future tutorial). Given the tile-type storage of the tiff pages, it makes it possible to load only the necessary tiles (in near real time), so that as the user zooms in, the necessary tiles are loaded and interpolated for them to see.
Great. So how can we use these images in Matlab.
Matlab natively supports mutli-page tiff reading if you simply provide an index like so:
- io=imread('TCGA-A1-A0SD-01Z-00-DX1.DB17BFA9-D951-42A8-91D2-F4C2EBC6EB9F.svs','Index',2);
But this isn’t the whole story, using Matlab’s image info function:
- info=imfinfo('TCGA-A1-A0SD-01Z-00-DX1.DB17BFA9-D951-42A8-91D2-F4C2EBC6EB9F.svs');
we see that there are additional levels present in the tiff image.
What are the additional images?
Page 6 contains the ID, this is usually a patient number or some other type of identification number written (or printed) at the end of the slide.
Page 7 contains a view of the entire slide, which has been automatically cropped (in green box) to show only an area where there is material. Imagine having to store values for “everything”, even the white space of the microscope slide where there is no additional information to be had. So this page shows us where/what has been taken from a high-level view of the entire slide.
We can see Aperio knows about these, and uses them for their front end, but does not report them in the information panel. On the other hand, Matlab has access to all of the information available.
This makes it a bit more tricky as the number of layers reported by Aperio and Matlab don’t line up, but the convention as defined by Aperio is rather straightforward:
| Level | Content |
| First level | Full resolution image |
| Second level | Thumbnail |
| Third level to N-2 Level | A reduction by a power of 2 (4:1 ratio, 16:1 ratio, 32:1 ratio, etc) |
| N-1 Level | Slide Label |
| N Level | Entire Slide with cropped region delineated in green |
With this information at hand, we can decide which level we want to load and can at least start to do some work. The next part of this tutorial discusses how to load only specific sub-sections of the high-resolution image given lower-resolution information.





Thanks for being so helpful as a blogger and leader in the field.
I have few quick questions about the naming of the files.
1. TCGA-FG-8187-01Z-00-DX1.4af1e387-0e5f-43e2-a237-
fad93a0209a7
2. TCGA-FG-8187-01Z-00-DX2.e9ba4a17-f5b7-4786-b4f3-
29e0d79b5985
Questions;
a) Are those two files from same patient?
b) What these DX1 and DX2 mean?
c) I see some files are named with DX5 also. I just one to know exactly each part of the name of the slide images.
Thanks in advance
Reza
I see why this could be confusing!
Here are the answer to your questions:
a) yes, both files are from the same patient.
b) DX stands for diagnostic. There are thus sometimes X slides for a patient if the X-1 diagnostic slides don’t provide enough certainty in a diagnosis for the patient. I’ve seen a few instances where DX1 is a frozen specimen, of much lower quality, taken during the surgery so that the surgeon can gain immediate insight into the boundaries of the tumor. The DX2 is then done later in a laboratory using high quality paraffin fixing methods.
c) DX5..i guess they kept going until they got what they needed. Its important for them to continue labeling it, since i’m sure there are slides DX1…DX4 in the set somewhere as well (maybe not uploaded to TCGA), and they need to prevent a file name collision later on. Better safe than sorry!
Thanks for reading my blog!
Cheers,
Andrew
Dear Andrew,
Thanks a lot for reply. Your blog is really helpful for those who are doing research in these fields. I appreciate your works.
Best Regards,
Syed Reza
I read the .svs data in matlab. However, all of the values are zero and cannot process the image on Matlab. Could you tell me how to solve this problem.
The simple Matlab code is:
img=imread(‘TCGA-A1-A0SD-01Z-00-DX1.DB17BFA9-D951-42A8-91D2-F4C2EBC6EB9F.svs’,’Index’,1);
all of the values in variable “img” are zero.
Try loading the second index instead of the first one.
The first index is likely too big to fit into RAM:
94,075 (height) * 80,287 (width) * 3 (RGB) = 22,658,998,575 uint8 integers (or ~23GB)
i have the same problem did u resolve that?
i have a problem when access the highest resolution page in svs file which is one . the image is black
could u help me to resolve that?
You can try using the openslide library instead of the matlab imread function
openslide library is also producing the same results. all black images.
I got it working with openslide
fantastic! maybe you can share with others, so if the problem happens to them they can fix it? what did you do to get it to work?
is there anyway to have a quick installation of MATLAB?
Newbee here. Please help me
sorry, no there is not. one would have to purchase it
I currently have IMageJ in my unit but I don` t have much experience in using the platform but I did some research about it to somehow have knowledge with the basics adI found that it is much more effective but the problem that I have is the macro commands that will be used since I don` t have that knowledge with writing scripts. Could you help me do some script writing/ commands?
Thank you
sorry but unfortunately given my limited bandwidth I’m unable to provide assistance with individual projects. I can certainly recommend some consultants for you if you’re interested
Please do. I would be delighted for your help
I can highly recommend Rob Toth: https://www.robtoth.net/contact
I am a student currently working on .svs images .
and i am working in python , does anybody know how to label the data with different patch size
sorry, i don’t know what this means, can you be more precise?